Quick start
Chinese homepage link: http://phylosuite.jushengwu.com/
1. Installation
1.1 PhyloSuite
Please see http://phylosuite.jushengwu.com/dongzhang0725.github.io/installation/.
1.2 Plugins
Please see http://phylosuite.jushengwu.com/dongzhang0725.github.io/PhyloSuite-demo/how-to-configure-plugins/.
2. Operation
For the basic operation of PhyloSuite, please see http://phylosuite.jushengwu.com/dongzhang0725.github.io/documentation/#4-1-0-1-Brief-example.
3. Create a workplace
When starting a new project, it’s recommended to create a new workplace for it.
- Hovering over
WorkPlacemenu bar will popup available workplaces, selectOthersto open theSet workplacewindow; - Click
add buttonto select a workplace folder.
4. Create a work folder
Work folders are where you can deposit input data and results for different analyses.
- Click the green
addbutton; - Enter the new work folder name.

5. Import sequences
Please see five ways to import sequences into PhyloSuite.
6. Start the analysis
6.1. Examples and test run
Please see http://phylosuite.jushengwu.com/dongzhang0725.github.io/example/
6.2. Example of each function
You can access a brief example demo of each function via the question mark button in the window of the corresponding function. For example:
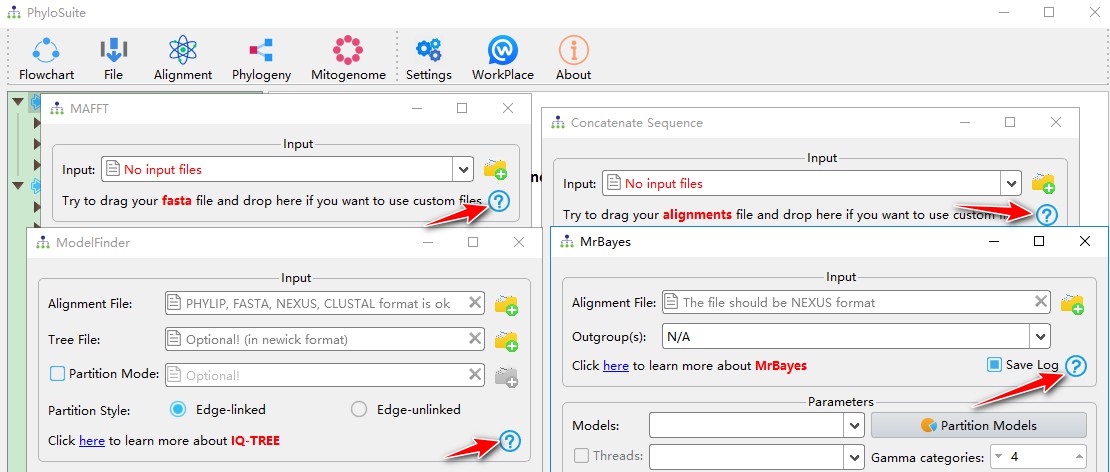
6.3. Comprehensive demo tutorials
For comprehensive tutorials for different analyses, please see single-gene tutorial and muti-gene tutorial.
7. Get the results
7.1. Select a work folder
By default, if you select a work folder or a results folder in a work folder before opening the function window, the results will be deposited in the selected work folder. Otherwise, the results will be deposited in GenBank_File/files or Other_File/files work folder. You can select a work folder via the ‘down arrow’ menu of the Start button in each function.
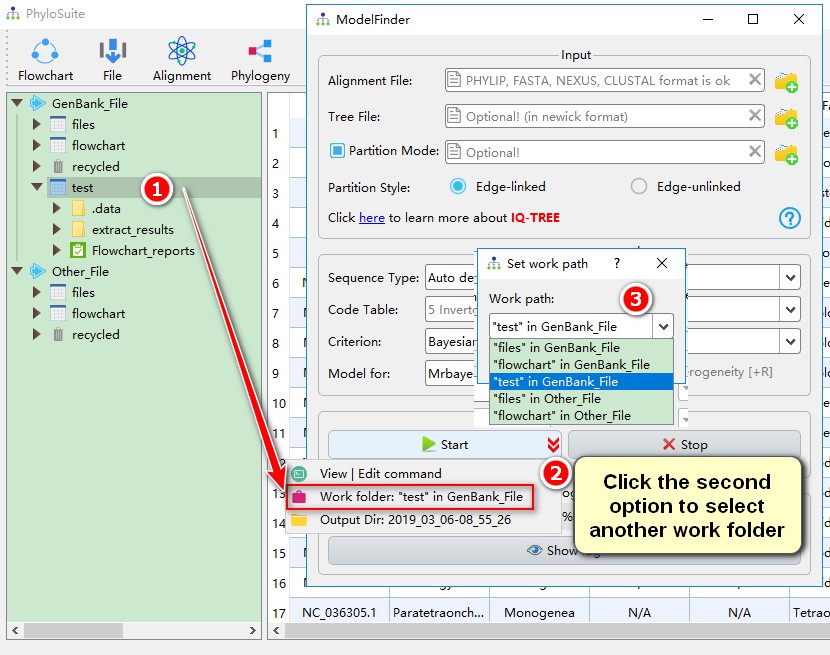
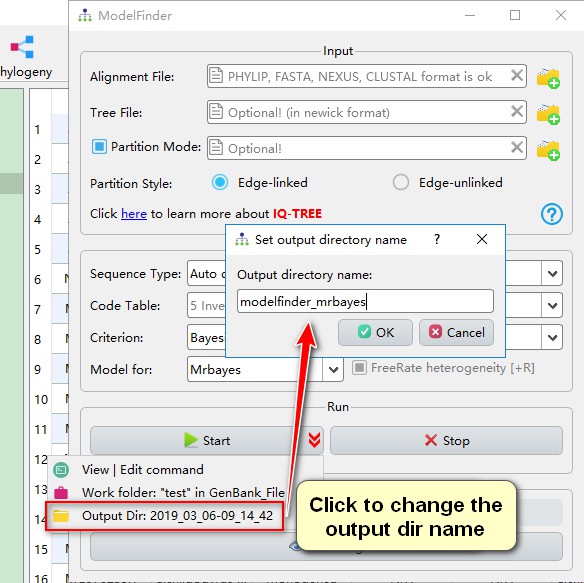
7.2. Set the output directory name
Starting with the version 1.1.15, PhyloSuite allows results of multiple analyses to be stored within each results folder in different output directories within a single Results folder. The default format of the directory name is: year_month_day-hour_minute_second, but you can change it according to your preferences.
7.3. Delete results
You can remove results you don’t need any more through the Remove function, accessed via the right-click menu.
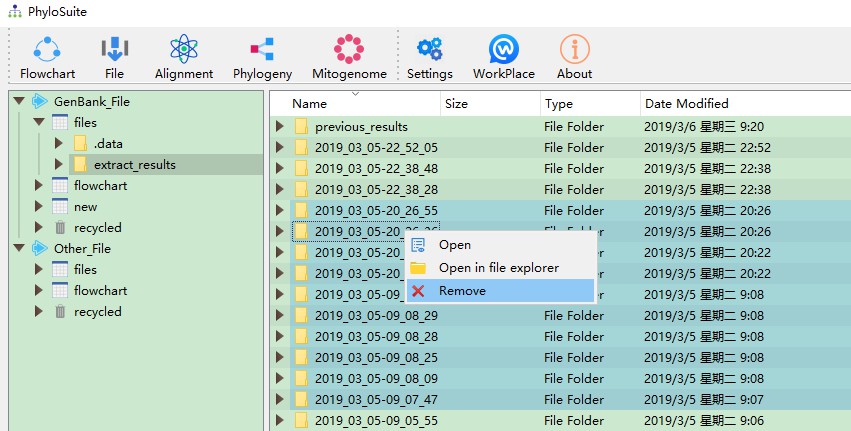
However, we recommend to remove them in your local file system, as that will enable you to recover them using the inbuilt operating system file recovery function. You can enter the local file system via the Open in file explorer option (see the figure above).
8. Recommended reading
Other demo tutorials can be seen here.